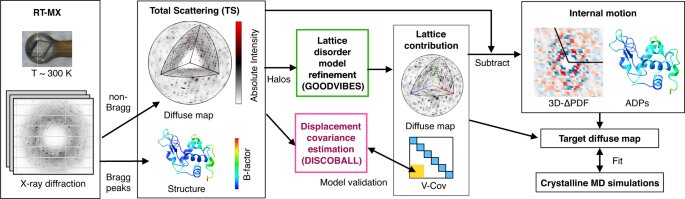
A novel two-part method for interpreting X-ray crystallography experiments on protein crystals has been developed by researchers at Cornell University. The method, outlined in a paper published on March 3 in Nature Communications, enables data, which was formerly discarded, to be analysed, leading to a better understanding of a protein’s movement, structure and function. Protein crystallography produces Bragg peaks, which provide high-resolution information about protein structure, as well as capturing cloudy images related to protein movements which are typically ignored. The new method separated these movements and produces a structure of the protein and its atomic movements. The goal is to create a new structural technique – GOODVIBES and DISCOBALL – which researchers can use to test their hypotheses and better scrutinise protein data from X-ray crystallography experiments.
More information: Steve P. Meisburger et al, Robust total X-ray scattering workflow to study correlated motion of proteins in crystals, Nature Communications (2023). DOI: 10.1038/s41467-023-36734-3
Denial of responsibility! TechCodex is an automatic aggregator of the all world’s media. In each content, the hyperlink to the primary source is specified. All trademarks belong to their rightful owners, and all materials to their authors. For any complaint, please reach us at – [email protected]. We will take necessory action within 24 hours.

Jessica Irvine is a tech enthusiast specializing in gadgets. From smart home devices to cutting-edge electronics, Jessica explores the world of consumer tech, offering readers comprehensive reviews, hands-on experiences, and expert insights into the coolest and most innovative gadgets on the market.