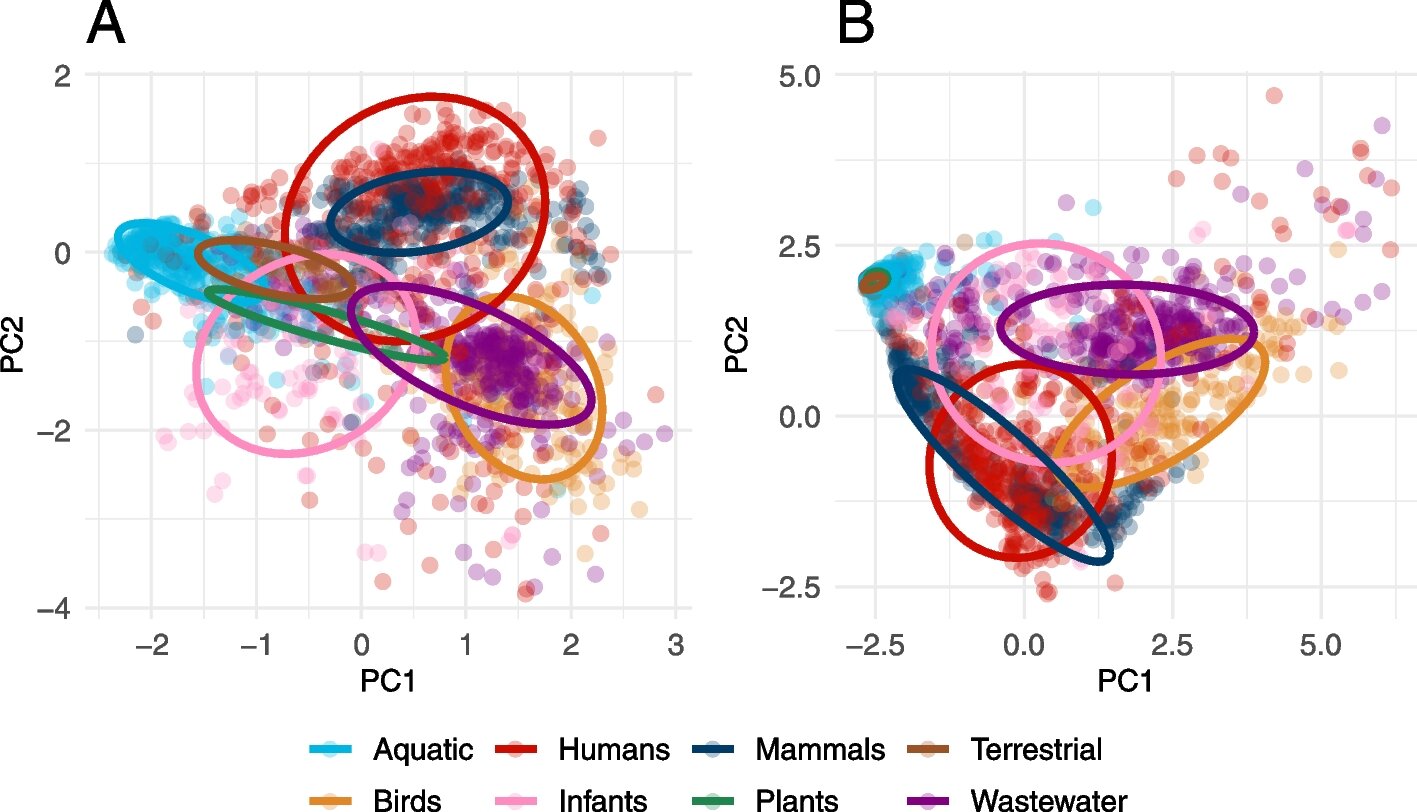
The abundance of latent and established antibiotic-resistant genes (ARGs) was analyzed using principal component analysis in various environments including aquatic, plants, infants, wastewater, terrestrial, human, birds, and mammals. The study, published in Microbiome by Swedish researchers from Chalmers University of Technology and the University of Gothenburg, found that ARGs are more prevalent in bacteria in all environments than previously thought. The study analyzed 630 billion DNA sequences, identifying new forms of resistance genes in various environments, such as sewage treatment plants, soil, and even the human microbiome. This presents a serious threat to global health, as antibiotic-resistant bacteria are difficult to treat. The researchers recommended a focus on enhancing understanding of the development of resistance in bacteria through risk assessments relating to antibiotic resistance.
Denial of responsibility! TechCodex is an automatic aggregator of the all world’s media. In each content, the hyperlink to the primary source is specified. All trademarks belong to their rightful owners, and all materials to their authors. For any complaint, please reach us at – [email protected]. We will take necessory action within 24 hours.

Jessica Irvine is a tech enthusiast specializing in gadgets. From smart home devices to cutting-edge electronics, Jessica explores the world of consumer tech, offering readers comprehensive reviews, hands-on experiences, and expert insights into the coolest and most innovative gadgets on the market.